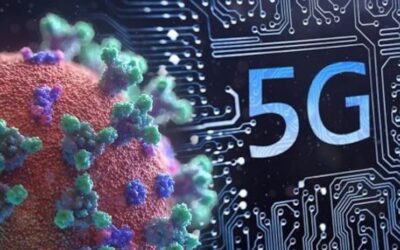
In June 2011, a virulent strain of E. coli swept across Europe, killing 22 people and sickening over 2,153 others with severe symptoms including bloody diarrhea and hemolytic uremic syndrome, a condition that causes kidney failure. The outbreak was eventually traced to a sprout farm in northern Germany, but the most alarming aspect of the crisis was not its origin but rather the pathogen’s extraordinary genetic profile.
The strain, identified as E. coli O104, displayed resistance to eight distinct classes of antibiotics, a characteristic so unusual that it prompted questions about whether such a microorganism could have developed through natural evolutionary processes alone.
Genetic Analysis of the O104 Strain Revealed Unprecedented Resistance
Researchers at Germany’s Robert Koch Institute sequenced the genome of the O104 strain and discovered it was impervious to an extraordinary range of antimicrobial drugs. The list of ineffective treatments included penicillins, tetracycline, nalidixic acid, trimethoprim-sulfamethoxazole, cephalosporins, amoxicillin combined with clavulanic acid, piperacillin-sulbactam, and piperacillin-tazobactam.
Beyond simple drug resistance, the strain also produced Extended-Spectrum Beta-Lactamases, commonly abbreviated as ESBLs. According to the UK Health Protection Agency, these enzymes enable bacteria to neutralize cephalosporins such as cefuroxime, cefotaxime, and ceftazidime, which represent some of the most commonly administered antibiotics in hospital settings worldwide.
The O104 variant also carried two particularly concerning genes designated TEM-1 and CTX-M-15. The Guardian reported that these genetic markers had alarmed infectious disease specialists since the 1990s due to their association with critical organ failure and high mortality rates in infected patients.
How Multi-Drug Resistance Typically Develops in Bacteria
Antibiotic resistance in bacteria generally emerges through a process of selective pressure. When a bacterial population is exposed to an antimicrobial agent, the vast majority of organisms die, but any individual bacteria carrying a random mutation that confers survival will reproduce and pass that resistance trait to subsequent generations.
Developing resistance to a single antibiotic through this mechanism is well documented and occurs regularly in clinical and agricultural settings. However, acquiring resistance to eight separate classes of drugs requires sequential exposure to each class, with surviving colonies being isolated and subjected to the next antimicrobial agent in succession.
This process of stepwise selection is, in fact, the standard methodology used in laboratory settings to create resistant bacterial strains for research purposes. Biodefense laboratories, including facilities such as the National Biodefense Analysis and Countermeasures Center at Fort Detrick, Maryland, maintain the capability to engineer precisely these types of multi-resistant organisms for defensive research and countermeasure development.
The Statistical Improbability of Natural Eight-Class Resistance
Standard O104 strains of E. coli are not typically resistant to antibiotics under normal conditions. The probability of a single bacterial strain independently developing resistance to eight different drug classes through random mutation in a natural environment is extraordinarily low from a genetic permutation standpoint.
Antibiotics are not used in vegetable cultivation, which further complicates the narrative that this superbug emerged naturally from the food supply chain. Without sustained antibiotic exposure, there would be no selective pressure to drive the development of drug resistance in bacteria colonizing produce.
This statistical argument formed the basis for speculation that the O104 strain may have been deliberately engineered in a laboratory setting and subsequently introduced into the food supply, either intentionally or through accidental release. Critics of this hypothesis countered that horizontal gene transfer between bacteria, the sharing of resistance genes through plasmids, could theoretically produce multi-drug resistant organisms without direct sequential antibiotic exposure, though achieving eight-class resistance through this mechanism alone would still be highly unusual.
The European Regulatory Landscape at the Time of the Outbreak
The timing of the E. coli crisis coincided with significant regulatory developments across the European Union. The EU had recently implemented restrictions on medicinal herbs and nutritional supplements under the Traditional Herbal Medicinal Products Directive, which went into full effect on April 30, 2011, just weeks before the outbreak began.
Some commentators drew connections between these regulatory actions and the food safety panic generated by the outbreak, suggesting that the crisis served to justify expanded governmental control over food production and distribution. Germany’s agricultural ministry notably declined to lift warnings against consuming tomatoes and lettuce even after the actual source of contamination had been identified as sprouts, a decision that prolonged economic damage to vegetable producers across the continent.
Spain’s agricultural sector was particularly affected after initial reports incorrectly identified Spanish cucumbers as the contamination source. Leaked diplomatic cables previously published by WikiLeaks had revealed tensions between Spain and the United States over Spain’s resistance to adopting genetically modified crops, adding another layer of geopolitical intrigue to the crisis narrative.
Public Health Impact and the Question of Treatment Alternatives
The outbreak ultimately affected ten European nations, with the majority of cases linked to individuals who had visited northern Germany. The combination of the O104 strain’s antibiotic resistance and its ability to cause hemolytic uremic syndrome made treatment exceptionally difficult using conventional pharmaceutical approaches.
The crisis highlighted the growing global threat posed by antibiotic-resistant bacteria, regardless of whether this particular strain emerged through natural processes or laboratory manipulation. The World Health Organization has repeatedly warned that antimicrobial resistance represents one of the most pressing public health challenges of the coming decades.
Research into alternative antimicrobial approaches, including plant-derived compounds from garlic, ginger, and various medicinal herbs, as well as probiotic therapies designed to restore healthy gut flora, gained renewed attention in the aftermath. These complementary approaches, while not replacements for conventional treatment in acute cases, represented an expanding area of scientific inquiry into non-pharmaceutical methods of managing resistant bacterial infections.
The 2011 European E. coli outbreak remains a landmark case study in infectious disease epidemiology, antibiotic resistance, food safety regulation, and the complex intersection of public health, agriculture, and geopolitics.